Structural Biochemistry/Carbohydrates/Virus
Some viruses enter their host cells by attaching to cell-surface carbohydrates. The influenza virus in particular, attaches to sialic acid residues on the terminai of the oligosaccharides present near the cell-surface glycoproteins and glycolipids.

Once attached, viruses inject their own genetic material and take over the cell's machinery to produce more viruses. The cell can undergo a lytic or lysogenic cycle. In the lytic cell, the cell is taken over, produces viruses with its own machinery and organelles, and then dies, releasing more viruses. In the lysogenic cycle, the cell does not die but instead replicates with viral DNA/RNA in its own genome. A common example of a lysogenic virus is lambda phage. However, lambda phage can also enter the lytic cycle. In this manner, some viruses can remain dormant for years at a time, activating under certain environmental conditions to begin replication under the lytic cycle.
There are several different types of viruses, including DNA, RNA, and retroviruses. DNA viruses usually replicate by taking over the cell's DNA, whereas RNA viruses tend to replicate in the cytoplasm. Retroviruses transcribe their DNA into host DNA via reverse transcription- an example of this is HIV.
The body has several defense mechanisms against viruses. The body's first line of defense is the innate immune response, which is non-specific and defends the body against a variety of threats. Inflammation, coughing, sneezing, and a variety of other non-specific defense mechanisms are examples of the innate immune response in action. The adaptive immune system, on the other hand, targets specific threats. T Cell and antibody activation are examples of the adaptive immune system's defense mechanisms. In addition, scientists have developed vaccines and anti-viral drugs to assist the immune system and/or disrupt viral replication mechanisms.
Lymphocytes, or white blood cells, are the body's main defenders against foreign invaders. There are two major types of lymphocytes, T cells and B cells. Of the T cells, there are T helper, cytotoxic, memory, regulatory, and natural killer cells. Each type of T cell has its own purpose in immune response. Helper cells assist and activate other cells at the first signs of infection, cytotoxic T cells destroy infected cells, preventing more viruses from being released, memory cells store antigens, allowing the body to quickly recognize previous infections to make fighting future infections more efficient, regulatory T cells shut down immune response after an infection has subsided, and natural killer cells can help kill tumor cells. B cells, on the other hand, recognize antigens and develop anti-bodies once the correct antigen for an infection has been found.
Viruses are parasites because they can't survive on their own. They infect a cell and use the infected cell to multiply and make more viruses. Viruses are considered alive because they are capable of duplication and have defensive mechanisms. All viruses have some sort of genome, which can be single or double stranded, DNA or RNA, and linear or circular. A virus genome is also very small: they could be four genes or a few hundred when prokaryotes tend to have several thousands of genes and eukaryotes tend to have tens of thousands of genes. Viruses have a protein coat that protects the genome, and this coat is made up of protein. Usually, the coat has multiple copies of one protein because the virus wants to minimize its genome. Viruses want to pack up and ship off new copies of viruses as soon as possible, so if the genome is longer and more varied, then the longer the packing will take because it will have to encode for several different genes; thus, viruses would be at a great disadvantage if they had variety in their genome.
What is A Virus?
editA virus is an infectious, parasitic agent that can only replicate inside other host cells. All viruses have a genome and a capsid.
A viral genome can be:
•Single stranded (ss) or double stranded (ds)
•DNA or RNA
•Linear or circular
Compared to others, the viral genome is very small. It consists of two to several hundred genes. Prokaryotes, like bacteria, have thousands of genes, while eukaryotes can have tens of thousands of genes.
Viruses were once thought to have been a self-replicating genetic element but the guiding principle which differentiates the two is that “once a replicon incorporates a gene(s) that allows it to make a capsid to enclose the replicon, then a functional and structural entity called a virion is produced.” This means that the hallmark of a functional virus is a virion. [9]
The capsid is a protein coat that surround the genome. It consists of several copies of a single protein, which is advantageous for the virus because it only needs one gene to code for the capsid protein. The capsid and it's enclosed genome is also referred to as a nucleocapsid.
Some viruses also have a membrane bilayer around the nucleocapsid that can serve as a protective layer. This can be found in viruses that have already infected a host cell, and have budded off from the host cell.
Viral proteins have structures that are different from eukaryotic proteins in that they are loosely packed due to relatively fewer hydrogen bonding and van der Waals interactions. [2]
Viral proteins are small proteins with many disordered segments and many coil residues. The coil residues cause the protein to be loosely packed because the coil conformation (a secondary structure) is unorganized and does not easily lie near another protein strand. [2]
In addition, viruses are not considered living organisms as they fail to fulfill the basic requirements of life to be considered living. Although viruses replicate, they are not able to metabolize food. Viruses do not have an organized cell structure, it does not respond to any external stimuli when placed under such circumstances, nor do they maintain homeostasis in which internal temperature is attempted to be kept at a constant. Additionally, viruses do not belong to any animal kingdom, they are unable to adapt to environments, and their only method of reproduction is through invading host cells, but not being able to reproduce independently.
Origin of Viruses
editThe exact origin of viruses has been questioned by scientists for decades. Although there are many theories on the origin, there are three hypotheses that have been discussed at greater lengths about the origin of viruses.
1. Viruses originated from ancient times before cellular life was invented This hypothesis has been rejected many based on the idea that viruses need a cellular host in order to survive. If viruses existed before cellular life existed then they would not have had the ability to replicate in a host cell.
2. Viruses originated from cells by reduction. This hypothesis is rejected on the premise that this method requires intermediates between cells and viruses which have yet to be found.
3. Viruses escaped from cells by utilizing cellular replication elements removed from cellular control. This hypothesis doesn't explain how it is possible to build complex virion structures. Also since it has been found that viruses are capable to infecting cells in all three domains, it has been proven that they are ancient and not from reduced cells. Also the discovery of the mimivirus somewhat refutes this hypothesis.
When comparing a two different plant viruses, scientists were able to see the similarity in capsid structure and similarities in the arrangement within the capsid. When comparing two animal viruses, they were not only found to be similar to each other but also to plant viruses based on their coat protein folds. They all shared the stranded beta barrel folding. Plant, insect, and human viruses showed to share similar structural architecture which therefore led to the conclusion that ssRNA viruses may share a common ancestor. In addition to the ssRNA, after more comparisons, scientists saw that even dsDNA and bacterial viruses shared the same basic architect as ssRNA viruses; they all used the beta barrel fold. By comparing various viruses, it has been discovered that a common ancestor infected cells of all three domains before diversification. [9]
Virus Classification
editViruses are classified under two different classifications: the Baltimore Scheme and the International Committee on Taxonomy of Virus(ICTV).
The Baltimore Scheme divides all the various viruses into seven groups depending on their chemical type, number of strands, and if their single strand is capable of directly undergoing translation. The seven categories of viruses are dsDNA, ssDNA, dsRNA, plus(sense) ssRNA, minus(antisense) ssRNA, ssRNA with DNA intermediate, and dsDNA with RNA intermediate.
The second classification categorizes viruses into order, family, genus, species. The organization of the virus lineages is based on the information on the type of host cell each virus infects. Although the ICTV try to organize the viruses to the best of their ability, it is still difficult to designate a majority of the families to orders. [9]
Viral Structures
editViral structures have been found to be the unifying component within the viral universe. Viruses have several set structures and do not deviate from these structures. They can be icosahedral, helical, a combination of the two, bottle-shaped, lemon-shaped, and spindle shaped. Some viruses are also pleomorphic; they have a lipid envelope.
For classification purposes, the capsid structure has been declared to be the crucial element. The process of replication and the genomic structure were considered as an element in classification but because of the lack of information they provide for identity, it has been ruled out. The capsid structure is the main structural component that is congruent to the identity of the virus. [9]
Viral Reproduction
editViruses are obligate (imposed by necessity; incapable of adaptation to different conditions; restricted to a particular mode of life) intracellular parasites. They reproduce only within a host cell. Viruses contain no metabolic enzymes or "machinery" for protein synthesis. They can only infect a limited range/type of host cell (some viruses can infect several species, such as the rabies virus; some viruses can only infect one species; and animal viruses are usually tissue or cell-type specific; that is they will only infect one particular type of cell). Steps of viral life cycle (viral binds to host cell; lock and key fit between viral protein and host cell surface receptor; viral genome, via variety of mechanisms, enters cell; viral genome 'commandeers' its host, using host cells machinery to copy viral genome and synthesize viral proteins; DNA viruses usually use the host cell's DNA polymerase-such as reverse transcriptase).
Animal Viruses
editAnimal Viruses are very diverse with many modes of reproduction. Many animal viruses have outer membrane (viral envelopes). Typically there is a lipid bilayer (derived from the host cell plasma membrane) with virally encoded proteins protruding from it. These virally encoded proteins are important for binding and helps virus enter the cell.
Herpes virus envelop derive from nuclear envelope (herpes virus genome integrated into host cell DNA - as provirus). Usually the virus remains latent. But stress causes virus to become active. The provirus leaves genome and initiates viral production. Blisters form as a result.
DNA Viruses
editThere are two types of DNA viruses: double stranded DNA viruses and single stranded DNA viruses.
•Double stranded DNA viruses, such as the Small Pox virus, are able to use DNA polymerase to replicate itself, RNA polymerase to transcribe mRNA, wherein it will then use the host cell's ribosomes to produce proteins.
•Single stranded DNA viruses will have to use DNA polymerase twice to replicate its own genome (first synthesized strand will be a conjugated gene expression of the viral genome), but they can still use RNA polymerase to produce mRNA and the host cell's ribosomes to produce proteins.
RNA Viruses
editThere are three types of RNA viruses: double stranded RNA viruses, single stranded RNA viruses, and the retrovirus. Unlike DNA viruses, RNA viruses can use RNA replicase to replicate in a host.
•In a double stranded RNA viruses, each strand is considered to be viral mRNA. RNA replicase can be used to both duplicate the genome and produce viral proteins.
•There further exist two types of single stranded RNA viruses, positive and negative. In positive SS RNA viruses, the RNA is known as a "sense" strand, and acts as the mRNA. RNA replicase will make a complement of the mRNA, which can then act as a template to replicate it's genome. Positive SS RNA viruses also use RNA replicase to translate proteins. In negative SS RNA viruses, their genome is considered a "nonsense" strand, and is not the same as the mRNA strand. As a result, RNA replicase must make a template strand off of the "nonsense" strand and this can then be used to produce proteins. It also serves as a template to replicate new, "nonsense" RNA.
•Retroviruses are also single stranded RNA viruses, however, their genome is converted into double stranded DNA through viral reverse transcriptase. First, Reverse transcriptase will make a DNA/RNA hybrid strand using the original single stranded RNA genome. Then it will use the newly synthesized single stranded DNA template to make a double stranded DNA, and then integrate it into the host genome where it will be duplicated during the host cell's cell cycle.
Viroids and Prions
editViroids are small molecules of naked RNA that infect plants. Prions are infectious proteins (cause a number of degenerative brain disease). Brain diseases: mad cow diseases, scrapie in sheep, and Creutzfeldt-Jakob disease in humans. It may be a misfolded version of a normal protein, which redirects protein folding in infected cells. This generates more misfolded protein which can infect other cells.
Fighting Viruses
editHost Cell Defenses
Viruses are often hard to see in cells because their "envelope" makes them look like parts of the host cell. However, if the cell sees double stranded RNA, it alerts the cell that something is wrong. Double stranded RNA is considered to be foreign, and once it is spotted, dicer enzyme will cleave it into smaller RNA pieces. RNA interference (RNAi) will come and destroy the small double stranded RNA pieces. Double stranded RNA will also signal transcription to stop, and increase the amount of ribonucleases, an enzyme that catalyzes the degradation of RNA. This will get rid of all the double stranded RNA as well. Once all the double stranded RNA is destroyed, transcription will continue as normal.
If a virus inhibits dicer and the cell can't eliminate the virus inside the cell, the infected host cell will be tagged for apoptosis.
An immune response through the Major Histocompatibility Complex I (MHC I) occurs as well. MHC I are involved in antigen presentation. Once the host cell cleaves the virus into smaller pieces, MHC I can present the antigen outside the cell, which can then be recognized by T cells and promote antibody production to help kill the infected cells.
Treatments
Viruses have structures that easily change conformation to adapt to new hosts and conditions, which explain how they elude drug treatments.[2] For example, there are new flu shots available every year to combat new strains of the flu virus, which arise from minor mutations that allow them to better exist in certain conditions.
Antibiotics do not work on viruses, instead, vaccines are synthesized invitro. Vaccines are made through purified viral proteins. The purified protein, lacking the virulent genomic information, can be cleaved into pieces and injected into a host. This way, the host will not risk the virus infecting its cells, rather, the body will recognize the cleaved pieces as foreign antigen from the virus and mount an immune response against it. This enables the immune system to have antibodies prepared and be ready in the event of a real infection.
Reverse transcriptase inhibitors (RTIs) are a type of antiretroviral drugs, that target RNA viruses such as HIV infections. RTI is used to inhibit the activity of reverse transcriptase which is a required enzyme that allows the reproduction of viruses. By inhibiting transcription, viral DNA is unable to be transcribed from RNA, which halts the viral infection cycle. However, with time HIV infected cells will eventually mutate into another form with a different kind of reverse transcriptase tag unaffected by antiretroviral drugs usually across a span of about five years.
Flu shots usually contain 3 different types of inactivated viruses, because researchers cannot always predict the exact type of virus that plays the biggest role in the seasonal flu. They hypothesize the protein structures of the 3 that have the highest probability and put those into the vaccine.
Resistance
Many antiviral agents are inhibitors that appear like the actual molecule that the virus' proteins want to bind to. By taking the place of the actual protein, the inhibitor is able to act like a "decoy" and block up the active site of the virus' protein and help stop the virus' life cycle.
Because the virus' protein's active site cannot easily be changed, the effectiveness of the inhibitor being used depends on how well it fits into the active site. Thus, oftentimes, the closer that the inhibitor's structure is to the actual molecule that usually fits into the active site, the more effective the inhibitor. It has been found through clinical tests that the closer the inhibitor's structure is to the actual substrate, the higher the barrier to resistance by the virus.
Viruses also have machinery that allows them to fuse with host cells. This fusion machinery has also become a primary target of drugs. Viruses such as HIV have shown that membrane fusion follows the formation of a six-helix bundle. This bundle is created through a pathway in which there is a trimeric coiled-coil intermediate which can associate with the target membrane by use of a "fusion peptide" at the N-terminal end.
Large problems with viral resistance have arisen. Higher than normal rates of viral resistance within a population can often be attributed to patient use of the antiviral agent. By not taking the proper dose or not using the drug for the prescribed duration of time helps lead to viral resistance. This is because without the proper dosage of the drug (i.e.: less taken than prescribed), not enough of the inhibitor is present to take the place of the actual substrate. Thus, strains of the virus that are only mildly resistant to the drug and would normally die under the prescribed conditions will end up surviving and passing on its resistance genes to its progeny. After a certain amount of time, this weak resistance is able to become strong enough after multiple mutations to fight off the antiviral agent, even under prescribed conditions. Through the selection process of non-resistant strains being removed from the genepool, more and more strains are becoming resistant.
A large problem with anti-viral resistance is the responsibility that everyone in a given population has to help prevent its spread. One person's negligence in not taking his or hers prescription properly can help the resistant strain spread - giving other people less of a fighting chance when they themselves become infected. Unfortunately, even with the utmost diligence in following a given prescription, viruses can still become resistant after a certain amount of time. So even though resistant viral strains are ultimately becoming more and more common no matter what we do, people should still take personal responsibility to wholeheartedly follow a given regiment to help slow the spread of this phenomenon.
Viral Benefits
editThere are a few benefits that viruses provide for humans.
•Viruses can help us to understand our cellular machinery.
•Viruses attack harmful insects and bacteria.
•Viruses also provide a hope for gene therapy. Gene therapy is the process of introducing foreign DNA into a cell. In theory, one can obtain an empty virus (one that's had its disease-causing genome removed), load it with a specific DNA, amplify the virus through PCR to have a specific protein that can target receptors on a specific cell, and have it inject the DNA into the cell. For example, cystic fibrosis occurs due to the lack of CFTR, so in theory, one can load an empty virus with CFTR DNA, tag it with something that's specific for lung receptors, and inject it into a host. However, in reality this is very difficult to do. Furthermore, viruses have a relatively high transfection rate. In other words, they are able to successfully transmit their genetic material into the host cell at a high rate and with less complications than more traditional methods that often cause unnecessary cell death.
Influenza
editThe influenza virus is a negative single stranded RNA virus. 2 types pose the greatest threats to humans: type A and type B. It has eight strands of RNA that encode for eleven different genes. These genes are then translated into many different proteins, but two major types of viral proteins are:
•Hemagglutinin (HA), 16 different antigens exist Hemagglutinin is found on the surface of the viral protein and binds to the sugars.
•Neuraminidase (NA), 9 different antigens exist Neuraminidase (sialidase) cleaves the oligosaccharide chains which releases the virus' progeny to infect other cells. This viral protein has been targeted for anti-Influenza treatment. Neuraminidase inhibitors including Zanamivir and Oseltamivir (2 anti-Influenza drugs) are different structural analogs of the substrate that attempt to fit into the virus' neuraminidase binding site in order to prevent it from attaching to the actual cellular receptor. 10 distinct forms of neuraminidase have been discovered - 9 of them called "N1, N2, N3, ..., N9" for Type A Influenza and "Type B neuraminidase" for Type B Influenza. Although Type A and B's neuraminidase catalytic machinery are essentially the same, differences in the amino acid sequence in other regions of the protein cause slightly different reactions of each towards various neuraminidase inhibitors.
Current research in neuraminidase inhibitors has found that many analogs of current inhibitors preferentially bind to Type A influenza over Type B, making them poor choices as inhibitors. These changes that make the inhibitor select preferentially for Type A include analogs of the carboxamide chain in Neu5Ac2en.
The N1, N4, and N8 subtypes of neuraminidase in Type A Influenza are special, relative to the other subtypes because they contain an additional subsite within the binding site. The amino acid sequence responsible for this has not yet been identified. Furthermore, no biological function has been found to be associated with this additional subsite.
HA is a viral protein that binds to the host cell for entry. It specifies which cells can be infected. NA is the viral protein that is responsible for viral release by cleaving the HA/host cell interaction.
Influenza strains consist of different combinations of HA and NA, as well as other viral proteins.
A growing concern among many is the lethality behind the different strains of influenza. For example, the Bird Flu, Swine Flu, and the common flu.
•Bird Flu (H5N1): This strand does not infect humans easily and is not easily spread. There have only been 400 cases in the last 5 years, however, there were 262 deaths, giving H5N1 ~65% mortality rate (11/2009). H5N1 targets the lungs, and while it is not something many have to be concerned with at the present time, it will grow to be a very severe problem if the strand mutates into a form where transmission can occur from human to human.
•Swine Flu (H1N1): This strand, unlike H5N1, infects humans easily and can be spread easily. There have been 14-34 million estimated cases, with around 2,500-6,000 deaths, giving H1N1 less than a 0.01% mortality rate (11/2009). This shows that H1N1 is not very virulent, which is very good for humans. However, if the strand ever becomes able to mutate into a virulent form, it would become a very severe problem.
•Common Flu: The common flu is constantly mutating each year, and there is an estimate of 25-50 million cases a year with around 30,000-40,000 deaths.
Resistance
editWhen Neu5Ac2en analogs were cultured with neuraminidase inhibitor-resistant influenza viruses, off-target mutations in receptor-binding sites of hemagglutinin occurred. This revealed a balance between the efficiency of demolition by neuraminidase and hemagglutinin’s affinity for the receptor.
Experiments were done with the catalytic site variants of neuraminidase. There was a resistant virus chosen by one of the carboxamide analogs that had shown a mutation in one of the three argininyl residues, R292. That same mutation is a characteristic known to all neuraminidases. A loss of enzyme activity can be achieved by substitution to K292, but a large loss of binding potency to carboxamide analogs of Neu5Ac2en will also occur. E276 has been proven to change the conformation of carboxamides bound with R292K variant to allow the hydrophobic pocket to bind. The interaction of E276 and R224 creates the hydrophobic binding pocket, but the resistance comes from the R292K variant. The resistance is a hydrogen bond between E276 and K292 seen in the crystal structure of the unliganded R292K variant.
The less the inhibitor resembles the substrate the greater the loss of inhibitory potency directed at R292K N9 neuraminidase inhibitors including: Neu5Ac, Neu5Ac2en, 4-amino-Neu5Ac2en, zanamivir, Neu5Ac2en carboxamides, and oseltamivir carboxylate.This finding supports the rule that the more a drug similarly resembles the natural substrates/ligands, the more effectively they will be able to suppress a drug-resistant virus. A minimal decrease of inhibitory activity toward the mutant and biological activity of the mutant is required in order to suppress a drug-resistant virus. The drug-resistant virus must preserve its binding ability in order to process the natural substrates or the drug that resembles the substrates.
Even though these in-vitro experiments are useful and educational, drug therapy on infected patients can prove to be quite different from these studies. The different dosages and methods of intake can result in different levels of drug at the site of the infection. A good example of this is the comparison of zanamivir and oseltamivir.
Bacteriophage
editBacteriophages are viruses that infect the bacteria. A good example is the bacteriophage T2, which infects Escherichia coli. The T2 and T4 bacteriophages have a capsid with a needle like tail that can insert their own genome into the host cell, where it can instruct the reproduction progeny virions. The newly formed virions can then escape the host cell through a common phenomenon known as lysis. The location where the lysis occurred can be conspicuously observed through the formation of plaques, a clear spot surrounded by a plethora of bacterial cells. Plaques form by a single phage particle that lyse a host cell which then can infect nearby cells.
Bacteriophage life cycle
To begin its life cycle, the phage must find a host to attach to its surface. Cell surface receptor, a protein found on the surface of the host cell that is specific to the viral component, properly mediates the attachment and contact of the phage to the host.
The cell surface receptor plays an important role for the host cell, but the virus has mutated and evolved to take advantage these receptors. An example can be seen with the lambda phage. E. Coli has proteins called “lambda receptor proteins,” that allow the bacteria to acquire the sugar maltose for metabolism. However, the lambda phages have specific receptivity to the maltose porins in the outer membrane of E. Coli. Although this is detrimental for the host bacteria, natural selection has kept this porin for metabolic reasons.
With the insertion of the phage genome into the host cell, it directed the host to produce the progeny phages. Alfred Hershey and Margaret Chase In 1952 experimentally proved this when they have experimentally shown that when they transferred the DNA by a bacteriophage to a host cell, it led to the production of progeny phages. In 1950, Andre Lwoff and Antoinette Gutman observed that the phage genome can be integrated within the genome of the host bacteria. Most bacteriophages only insert their genome into the host through the cell envelope without the need for the whole capsid to penetrate the cell wall. T4 for example exhibit this kind of behavior. This virion has a neck tube that can contract and insert its DNA through the cell surface and into the host cell’s DNA.
Bacteriophage can go through two main cycles: The Lytic and Lysogeny cycle.
In a lytic cycle, the phage directs the immediate production of its progeny following the insertion of DNA. The process involves both the replicating the phage genome and the expression of the bacteriophage mRNA for the production of enzymes and capsid proteins. In some phages, like the T4, the host DNA is digested and increase the efficiency and of the bacteriophage production. After a plethora of progeny phage gets created, it proceeds to host cell lyses, which then releases the phages. Lysis is often referred to as a burst and the number of virus progenies released is called the burst size.
In the Lysogenic cycle, the phage inserts the DNA into the host cell but integrates its own genome into that of the host cell. Phage lambda, which has a linear genome of a double stranded DNA, reshapes the DNA to a circular shape upon entry into the host cell. The circular DNA can then integrate into the host genome by site-specific recombination of DNA. In this recombination, the recombinase enzyme aligns the Bacteriophage DNA with the host DNA so that the phosphodiester backbone links can be exchanged, which then leads to the integration.
The integration allows the phage genome to be replicated along with of the host cell as it replicates. The phage genome in the host DNA is called the prophage. Not only does Lysogeny integrate the phage genome into that of the hosts but also it can spontaneously generate a lytic burst of a phage. The prophage directs its own removal from the host genome by intramolecular process of site specific recombination with the two ends of the phage genome exchanging the phosphodiester backbone linkages once again. While the excised DNA exits the host genome, it circularizes and commences the lytic cycle, thus destroying the host cell and releasing the progeny phage.
Bateriophage can also go through a less prominent cycle called the Slow release cycle performed by the filamentous phages like the M13 phage. In this particular cycle, the phage particles replicate without the lysis of the host cell. The single stranded circular DNA of M13 serves as a template strand for the synthesis of a double stranded intermediate. Then this intermediate produces singe-stranded progeny genomes that get packaged by coating and supercoiling with the capsid proteins. These progeny phages force out through the host cell envelope without lysing the cell. The host cell continues to reproduce but more slowly because much of the resources are used to the production of the virus.
Proteins that bind DNA and subdue the transcription for the replication of the virus decide whether to go through the Lysis or Lysogeny. The transition from Lysogeny to Lysis can occur randomly but can also be affected by environmental factors such as UV light, which can damage the cell’s DNA. As for environmental cues, if a host cell’s growth is very strong, it’s more frequent to see the phage DNA inactive while an event that threatens the survival of the cell will initiate the lytic phage.
Virus transferring host genes
During the exit from the lysogeny phase, the phage can acquire the host genes and pass it onto another host cell in an event known as Transduction. Sometimes the whole phage genome can be permanently replaced by the host genome and packaged into the capsid, which would only be capable of transferring host DNA.
Lock and Key Mechanism
editAll viruses have a viral capsid. A viral capsid is the coat and the genome. However, only some viruses have envelopes. Envelopes are made up of lipids and they are essentially the same as the plasma membrane of the cell; they are similar because envelopes actually come from the host plasma membrane. Thus, the envelope has host and viral proteins that are gotten by exiting the host: viruses cannot do anything from inside the cell, so they have to get out to infect more cells. They do so by budding off and taking a piece of the host plasma membrane. The goal of a virus is to duplicate its genome and make protein; however, the real goal is to make more viruses, and the virus does so by infecting the host cell and working inside the host cell. First, the virus needs to figure out what sort of cells it can get into. The viral capsid and envelope proteins define the host/virus specificity: in the capsid there are specific proteins that will interact with specific proteins on the host cell membrane. This is the lock and key mechanism, where only certain viruses will recognize certain host cells. The host cell range is the types of cells that viruses can infect, and viruses have a limited host cell range: viruses can infect only specific cells and not all cells in the body, and the host cell range is define by the lock and key mechanism. The virus gets in the host cell by endocytosis (like the endosymbiont hypothesis) or by genome injection where only the genome is injected into the cell (the entire virus does not need to get in the cell since it only really needs the genome in). Both endocytosis and genome injection depend on the lock and key mechanism in order to decide which viruses can get in.
Lytic Life Cycle
editIn this cycle, the virus gets inside the host cell, duplicates its genome, makes protein, and then assembles new viruses by getting together the pieces it just produced. Eventually, the virus will cause the host cell to burst, which means that the lytic life cycle results in cell death and non-enveloped viruses. Everything in this cycle is done as quickly as possible, and the virus then moves on to other cells.
Lysogenic Life Cycle
editThis cycle does not result in any immediate killing of the cell. The virus enters the cell and actually inserts its genome into the host cell's genome. Then, when the host cell genome is duplicated, the viral genome gets duplicated along with it. Then, when something signals the virus to duplicate, the virus enters the lytic life cycle. The lysogenic life cycle includes the lytic life cycle.
Enveloped Virus
editThis type of virus can undergo either the lytic or lysogenic life cycle, but the end is different. After assembling, the viruses start budding out of the cell. They do not immediately kill off the cell but instead they take bits of the host cell plasma membrane. Enveloped viruses therefore contain many similar proteins and lipids to the host cells, which means that they are difficult to be detected by the immune system. Non-enveloped viruses, on the other hand, are easier to spot because they are seen as foreign. Also, some of the viral characteristics make them detectable as foreign: their double stranded RNA (our bodies do not have double stranded RNA) and viral proteins such as RNA replicase and reverse transcriptase (which are also foreign to the cell). Nonetheless, viruses are not as easily detectable as bacteria: while bacteria float around by themselves making them vulnerable to faster detection, viruses hide inside our body cells.
Genome packaging
editViruses use mainly two methods to package their genomes
- Building a capsid around viral genome
- Building a capsid first, then package genome into capsid; very often require the use of motor proteins
Packaging Initiation
editBefore packaging its own viral genome, there must be a process that can help viruses differentiate their own genome from its host genome; this process is known as initiation. There are several ways to initiate packaging:
- Unsegmented RNA
- Capsid has a binding site complementary to the specific sequence of the RNA or DNA
- Segmented RNA
- Segments have complementary sequence to other segments
- Capsid has more than one binding sites that recognizes the difference sequences on different segments
- DNA
- Double stranded DNA use a relatively different mechanism.
Examples of motor proteins and their mechanisms
editDifferent viruses have different types of genomes(dsDNA, dsRNA, ssDNA, ssRNA)and also different motor proteins. Not surprisingly, their packaging mechanisms differ from one another.
P4 ATPase
edit- Found in dsRNA viruses such as Φ6 and Φ12
- multi-subunit
- Has many other functions besides packaging genomes
P4 ATPase is a hexameric molecule that has a central channel that is lined with loops and helices[3]. Some of these loops have phosphate-binding sites. ATP will bind to these sites, resulting in a change in conformation within the central channel. This change in conformation is believed to be responsible for RNA translocation. P4 ATPase will remain in the final virion even when packaging is complete.
DNA motor proteins
edit- relatively powerful; most powerful known molecular motor is T4
- generates up to 60 picnewtons of force
- packages DNA at 700bp/s
Motor proteins that are used to package DNA genomes have to be more powerful than typical RNA motor proteins because DNA genomes have high density, which causes high pressures in the capsid(60 atm). Unlike P4 ATPase, DNA motor proteins will dissociate after completion of packaging.
Rotatory motor mechanism
editOne of the earliest motor protein component that was studied was the gp10 portal protein of Φ29. It was embedded within the capsid and its structure was determined to be similar to a funnal; the smaller end facing out of the capsid while the wider end faced the interior of the capsid. Its central channel was lined with α-helices that had negative charges on them[4]. This allows for the easy transition of DNA molecules since they are negatively charged themselves. It was predicted that the portal will rotate using energy from ATP hydrolysis, hence transporting DNA into the capsid. Cryo-electron microscopy discovered some structures on the capsid that supported this theory; however florescence spectroscopy experiments did not detect any rotation of the portal. This mechanism was therefore deemed unlikely.
Also, the shotgun method (also known as shotgun cloning) is a method in cloning genomic DNA. It involves taking the DNA to be cloned and cutting it either using a restriction enzyme or randomly using a physical method to smash the DNA into small pieces. These fragments are then taken together and cloned into a vector. The original DNA can be either genomic DNA (whole genome shotgun cloning) or a clone such as a YAC (yeast artificial chromosome) that contains a large piece of genomic DNA needing to be split into fragments.
If the DNA needs to be in a certain cloning vector, but the vector can only carry small amounts of DNA, then the shotgun method can be used. More commonly, the method is used to generate small fragments of DNA for sequencing. DNA sequence can be generated at about 600 bases at a time, so if a DNA fragment of about 1100kb is cloned, then it can be sequenced in two steps, with 600 bases from each end, and a hundred base overlap. The sequencing can always be primed with known sequence from the vector and so any prior knowledge of the sequence that has been cloned is not necessary. This approach of shotgun cloning followed by DNA sequencing from both ends of the vector is called shotgun sequencing.
HIV(Human immunodeficiency virus)
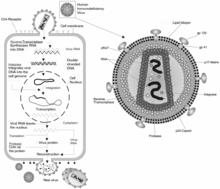
editThe HIV Virus is a kind of retroviridae called lentivirus. It infects vital cells in the human immune system and cause AIDS (acquired immunodeficiency syndrome) which will reduce the human immune system progressively. The HIV virus is difficult to fend off due to a few of its defense mechanisms including: Carbohydrate masking and the variance of its conformation.The patient will have a high risk of having life-threatening infections and cancers. HIV, just like other kind of Lentivirius, is transmitted as single-stranded, positive sense, enveloped RNA viruses. Unlike other retroviruses, HIV is roughly spherical with a diameter of 120 nm. It concedes of two copies of RNA that is positive single-stranded. The RNA is tightly bonded to nucleocapsid protein and the enzymes need for the development of the virus. There are two types of HIV, 1) HIV-1 and 2) HIV-2. HIV-1 is the majority of the HIV infection in the world since it is toxic and easier to infect other. It causes a progressive decrease of the CD4+T cell count. HIV-2 is infected per exposure. The HIV virus integrated into the host cell and become latent and cannot be detected by the immune system. There are 4 major ways to transfer HIV virus, unsafe sex, contaminated needles, breast milk, and transmission from mother to baby.
Some people, even if exposed to HIV, don’t develop AIDS because scientists discovered that they carried a rare genetic variant, which has slightly different sequence of nucleotides, that protects people from getting AIDS. It is called CCR5. This rare gene is thought to be selected during evolution because it made people resistant to an organism unrelated to HIV.
The time between getting the HIV virus in a human system and actually getting the disease associated with it (AIDS) is a very important factor to look into. During this time period, the human immune system gets progressively weaker, since the immune system is compromised. In fact, some viruses that people come into contact with (even the common cold) actually behave as cofactors to the HIV virus. However, to act as a cofactor, the other virus must have certain characteristics. First off, the other virus should be able to infect the same cells that HIV infects. Secondly, the amount of cells that get infected at the same time must be large enough to change the normal mechanism of the HIV virus. Many viruses have been proposed as cofactors to the HIV virus, but have failed to meet the first criterion listed.
Also, HIV viruses generally like to infect CD4+ T-cells. Many of these cells reside in the lymph nodes, so the HIV virus can also generally be found in the lymph nodes. HIV viruses can generally be found anywhere in the body that CD4+ cells are abundant. This includes places such as the adenoids, macrophages, and tonsils as well.
Many efforts have been made to create an effective vaccine against the HIV virus. Before being able to create the vaccine, the methods of transmission must be examined. This virus can be transmitted in four different ways. The most obvious of these ways is sexual transmission. The other paths of infection are needle sharing in drug users, mother to baby transmission, and the use of infected blood or products thereof. A good number of vaccines have been proposed for this, and are now in clinical trials. However, there are abundant ethical and social issues regarding the use of such vaccines on human volunteers. One concern is the obvious threat to human lives if the virus in the vaccine concoction gets out of hand. Another safety issue is that producing mass amounts of the retrovirus is hazardous to both the people working in the lab and the general public.
HIV Immunity
APOBEC is a protein that has been evolutionary conserved and is used in animals for making diverse proteins from mRNA. APOBEC3G is in the same family of conserved proteins, but is solely found in human beings. It is said to have an important job in anti-viral immunity, especially against retroviruses such as HIV, which is currently being studied. This symmetric protein, with 2 homologous catalytic sequences, is known to interfere with the reverse transcription activity of HIV before it can be integrated into the host chromosome. Typically, without APOBEC3G, a tRNA called tRNA3Lys binds to the HIV-1 primer binding site to start the process of reverse transcription. But when APOBEC3G is present, it can stop the primer binding site, which then stops the reverse transcriptase from making the single stranded DNA and eventually double stranded DNA.
However, there is something called the Viral Infectivity Factor (Vif), a protein that is native to the HIV virus, that is being researched because of its counteracting effects to APOBEC3G. Vif is known to attack APOBEC3G and deactivate it. Going back to how APOBEC3G functions, in the absence of Vif, APOBEC3G can catalyze dC to dU mutations in the reverse strand script, causing multiple copying errors in the daughter script. These are missense and nonsense codons that end up being copied.
Source: APOBEC3G: a Double Agent in Defense Harold C. Smith Department of Biochemistry and Biophysics and the Center for RNA Biology at the University of Rochester, School of Medicine and Dentistry, Rochester, NY 14642 USA
Reference
edit1. Berg, Jeremy "Biochemistry, 6th Edition" 2007
2. "Microbiology: an evolving science" by Joan L. Slonczewski and John W. Foster.
3. "Do viral proteins possess unique biophysical features?" by Nobuhiko Tokuriki1, Christopher J. Oldfield, Vladimir N. Uversky,
Igor N. Berezovsky, and Dan S. Tawfik.
4. "Biology" by Neil A. Campbell and Jane B. Reece
5. Colman, Peter M. "New Antivirals and Drug Resistance"
- ^ Sun, Siyang, Venigalla B. Rao and Nathan Nelson. “Genome Packaging in Viruses” Current Opinion in Structural Biology 20 (2010): 114-120. Pubmed. Web. 19 Nov. 2010.
Sun, Siyang; Rao, Venigalla B.; Rossman, Michael G. (2010), "Genome packaging in viruses", Current Opinion in Structural Biology, doi:10.1016/j.sbi.2009.12.006, PMID 20060706 {{citation}}: Unknown parameter |acess date= ignored (|access-date= suggested) (help)
Colman, Peter (2009), "New Antivirals and Drug Resistance", Annual Review of Biochemistry, PMID 19254207 {{citation}}: Unknown parameter |acess date= ignored (|access-date= suggested) (help)
6. The New Genetics - U.S Department of Health and Human Services
7. Colman, Peter M., “New Antivirals and Drug Resistance”, The Walter and Eliza Institute of Medical Research, 10.1146/annurev.biochem.78.082207.084029, March 2009, p. 95-112
8. Lever, A.M.L. "The Molecular Biology of HIV/AIDS". John Wiley & Sons. University of Cambridge Clinical School, UK. 1996.